We performed a meta-analysis to determine the effect Interleukin-6 (IL-6) promoter polymorphism (−174 G>C, −572 G>C, and −597 G>A) have on the development rheumatoid arthritis (RA) by ethnicity.
Material and methodsPubMed, EBSCO, LILACS, and Scopus databases were searched for studies exploring the association between any IL6 polymorphisms and RA until November 2018. Genotype distributions were extracted and, depending on the level heterogeneity, determined by the ψ2-based Q test and the Inconsistency Index (I2), fixed-effects or random-effects models were used to calculate pooled odds ratios (ORs) with 95% confidence intervals (95%CIs) for the heterozygous, homozygous, dominant, recessive, and allelic genetic models.
ResultsFrom 708 identified publications, 33 were used in this analysis. For the −174 polymorphism, Asians (ORheterozygous=7.57, 95%CI: 2.28–25.14, ORhomozygous=5.84, 95%CI: 2.06–16.56, ORdominant=7.21, 95%CI: 2.30–22.63, ORrecessive=5.04, 95%CI: 1.78–14.28, ORallelic=6.60, 95%CI: 2.26–19.28, p<.05) and Middle East countries (ORheterozygous=2.30, 95%CI: 1.10–4.81, ORdominant=2.27, 95%CI: 1.22–4.22, ORallelic=2.29, 95%CI: 1.24–4.23, p<.05) were associated with a significant risk of developing RA. Whereas, for Latinos, the C-allele was associated with a benefit (ORhomozygous=0.26, 95%CI: .08–.82, ORrecessive=.25, 95%CI: .08–.80, p<.05). For the −572 polymorphism, Asians demonstrated a significant association for the homozygous and recessive genetic models (8 studies, ORhomozygous=1.56, 95%CI: 1.16–2.09, ORrecessive=1.63, 95%CI: 1.08–2.45, p<.05). For the −597 polymorphism, no association was observed.
ConclusionsHere, the −174 G>C polymorphism increased the risk of developing RA in Asians and Middle East populations. Interestingly, for Latinos, the polymorphism was associated with a benefit. For the −572 polymorphism, only the Asian population showed an increased risk of developing RA for the CC genotype.
Realizamos un meta-análisis para determinar el efecto de los polimorfismos del promotor de interleucina-6 (IL-6) (-174 G>C, -572 G>C, y -597 G>A) sobre el desarrollo de artritis reumatoide (RA) analizado por etnicidad.
Materiales y métodosEn las bases de datos PubMed, EBSCO, LILACS y Scopus se buscaron estudios con la asociación entre polimorfismo de IL-6 y RA publicados hasta noviembre 2018. se obtuvieron las distribuciones de genotipo y de acuerdo al nivel de heterogeneidad el efecto fijo o aleatorio fueron utilizados para calcular los Odds Ratio (OR) con intervalos de confianza del 95% para los modelos genéticos heterocigoto, homocigoto, dominante, recesivo y alélico.
ResultadosDe 708 estudios identificados, 33 fueron utilizados para este análisis. Para el polimorfismo -174, los países Asiáticos (ORheterocigoto=7,57, 95%CI: 2,28–25,14, ORhomocigoto=5,84, 95%CI: 2,06-16,56, ORdominante=7,21, 95%CI: 2,30-22,63, ORrecesivo=5,04, 95%CI: 1,78-14,28, ORalélico=6,60, 95%CI: 2,26-19,28, p<0,05) y del Medio Oriente (ORheterocigoto=2,30, 95%CI: 1,10-4,81, ORdominante=2,27, 95%CI: 1,22-4,22, ORalélico=2,29, 95%CI: 1,24-4,23, p<0,05) están asociados con el riesgo de desarrollar RA significativamente. Mientras que, para los Latinos, el alelo-C está asociado con un beneficio (ORhomocigoto=0,26, 95%CI: 0,08-0,82, ORrecesivo=0,25, 95%CI: 0,08-0,80, p<0,05). Para el polimorfismo -572, los Asiáticos están asociados significativamente con los modelos genéticos homocigoto y recesivo (8 estudios, ORhomocigoto=1,56, 95%CI: 1,16-2,09, ORrecesivo=1,63, 95%CI: 1,08-2,45, p<0,05). Para el polimorfismo -597, no se observó asociación.
ConclusionesEl polimorfismo -174 G>C aumenta el riesgo de desarrollar RA en población Asiática y Medio Oriente. Curiosamente, para los Latinos el polimorfismo está asociado con un beneficio. Para el polimorfismo -572, solo la población Asiática demuestra una aumento en el riesgo de desarrollar RA con el genotipo CC.
The world prevalence of rheumatoid arthritis (RA) ranges between 0.5% and 1.0% of the adult population, in which women are four times more likely to develop the disease.1 RA is a chronic autoimmune disease, characterized by the inflammation of the synovium.2 RA can lead to destruction of the patient's joints and prolong/untreated RA can result in multiple organ manifestations, leading to severe disability and even mortality.3 Since RA is a multi-faceted disease, the exact etiology remains elusive. Evidence supports IL-6 as a factor in RA development, such as elevated IL-6 levels are found in RA patients’ serum and synovial fluid4,5; moreover, tocilizumab ameliorates disease activity and radiological progression.6 For a complete explanation of IL-6's role in RA, please see the review by Srirangan and Choy.7
IL-6 is a pro-inflammatory cytokine, released from many types of cells and is a mediator of the acute phase response.7 To date, many polymorphisms of the IL6 gene have been identified; however, the promoter polymorphisms have been shown to affect RA development.8–12 Three polymorphisms, −174 G>C (rs1800795), −572 G>C (rs1800796), and −597G>A (rs1800797) have been shown to augment or decrease IL-6 serum levels.13,14 Previous meta-analyses have shown that for the Asian population, the −174 G>C polymorphism increases the risk of developing RA15–17; however, for the −572 G>C and −597 G>A polymorphism, the results remain inconclusive. Moreover, with previous meta-analyses, there is an incomplete analysis with respect to Latin America and Middle East countries. Since the latest meta-analyses in 2016 for the −174 polymorphism, which included 13 to 15 studies, there have been 6 additional studies.9,18–23 Moreover, it is possible that between 7 and 10 studies were not included with these meta-analyses. Here, we preformed this meta-analysis to elucidate the effect the IL6 promoter polymorphisms have on the development of RA by ethnicity.
MethodsSearch strategyPubMed, SCOPUS, EBSCO and LILACS databases were searched for all studies that investigated for any IL6 polymorphisms and RA. The following keywords/index terms and any of their derivations were used: “Interleukin 6 or IL-6 or IFNB2”, “rheumatoid”, and “variant or SNP or polymorphism or genotype”. The search was performed without any language restrictions for publications published until December 15, 2018. Afterwards, the complied publications references were hand searched.
Inclusion and exclusion criteriaTwo authors independently determined if each study was to be included. If a disagreement occurred about a publication, a third author analyzed the publication in question. Initially, the titles and abstract were examined to determine if the article focused on RA and IL-6. Afterwards, the publications were thoroughly examined for the IL6 polymorphisms and the genotype distribution. For inclusion, the studies must have met the following criteria: (1) case-controls studies; (2) examined for any IL6 polymorphisms; (3) focused on human subjects; (4) RA-confirmed patients using either the American College of Rheumatology or the European League Against Rheumatism24 criteria; and (5) contained information about genotype frequencies. Studies were excluded if they were: (1) not a case-control study; (2) information was used in a previous publication; or (3) meta-analysis, reviews, or editorial articles.
Bias analysis and data extractionTwo authors independently assessed the quality of the studies using the Newcastle–Ottawa Quality Assessment Scale.25 The following aspects of each study were appraised: selection of cases and controls, comparability, and outcome or exposure. For the analysis, the possible quality scores ranged from 0 to 9 (see Supplement information). Studies that scored≥6 were considered as a high-quality study. The following data was collected from each study: first author's name, year of publication, geographical location, diagnosis criteria of RA, technique used to detect the polymorphism, source of controls, and genotype distribution for cases and controls. Before a study was to be excluded for missing information, we attempted to contact the corresponding author by email at least three times.
Statistical analysisFor each study, the Hardy–Weinberg Equilibrium (HWE) was determined by the ψ2-test for the controls and was considered in agreement when the p-value was>0.05. The crude odds ratios (ORs) and 95% Confidence Intervals (95% CI) were used to assess the strength of the association between the IL6 polymorphisms and the risk of RA. The pooled crude ORs were calculated for allelic (2 vs. 1), dominant (12+22 vs. 11), recessive (22 vs. 12+11), heterozygous (12 vs. 11), and homozygous (22 vs. 11) genetic models, where for the −174 polymorphism 1=G (Wild-type) and 2=C (mutant), for the −572 polymorphism 1=G (Wild-type) and 2=C (mutant), and for the −597 polymorphism 1=G (Wild-type) and 2=A (mutant). Heterogeneity was determined using the ψ2-based Q-test and its degree was assessed by the Inconsistency Index (I2). If there was not significant heterogeneity (ψ2-based Q-test p-value≥0.10 and I2<50%), the fixed-effects model was used (Mantel–Haenszel method), or if there was significant heterogeneity (ψ2-based Q-test p-value<0.10 and I2≥50%), the random-effects model was used (DerSimonian and Laird method) to calculate the pooled OR and 95%CI. Sensitivity analysis, removing one study and recalculation of the pooled OR, was conducted to verify the stability of the results. Begg's funnel plot, Begg–Mazumdar's test, and Egger's linear regression test were used to assess for publication bias. For geographic sub-analysis, initially, the studies were categories based on their country into Northern Africa, Sub-Saharan Africa, Latin America and the Caribbean, Northern America, Central Asia, Eastern Asia, South-eastern Asia, Southern Asia, Western Asia, Eastern Europe, Northern Europe, Southern Europe, Western Europe, or Oceania, according to United Nations M49 standard.26 Afterwards, the regions were group into Eastern Europe, Western Europe (Northern, Southern, and Western Europe), Latin America, Asian (Eastern and South-eastern Asia), and Middle East (Northern Africa, Southern and Western Asia). The Middle East category was based on the criteria that most countries identify as Arab and/or Muslim.27 All statistical analyses were conducted by using Comprehensive Meta-analysis v2 (Biostat, Inc., Englewood, New Jersey, USA). Unless noted otherwise, p-values <0.05 (two-sided) were considered statistically significant.
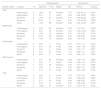
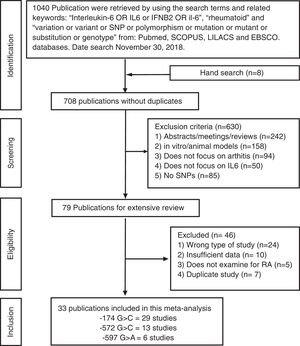
ResultsSelection of eligible studiesAfter duplicate removal, our literature search resulted in the recovery of 708 publications that focused on the association between the IL6 polymorphisms and patients with arthritis (Fig. 1). To note, 7 of the hand search articles were identified by the reference list, but the titles were originally in Chinese. After reviewing titles and abstract, 79 publications were considered for full-article examination. Of these publications, 24 were not case-control studies, 10 lacked sufficient data for analysis, 5 focused on the wrong pathology, and 7 were duplicate studies. Therefore, this meta-analysis contains 33 publications, which were divided between the −174 polymorphism (29 studies: cases=5920 and controls=6246), the −572 polymorphism (13 studies: cases=1973 and controls=1963), and the −597 polymorphism (6 studies: cases=1278 and controls=1524). The characteristics of the studies are presented in Table 1. 27.3% of the studies are from Asians countries: China,10,11,17,28–31 Taiwan,32 Japan.33 18.2% are from Eastern European countries: Russia,20,23 Czech Republic,34 Macedonia,35 Poland.18,36 24.2% are from Western European countries: Germany,37 Netherlands,12 Spain,38–40 Sweden41 and UK.42,43 21.2% are from the Middle East: Egypt,9,14,19 Iraq,22 India,15,44 and Turkey.8 Lastly, 3 studies are from Latin America.21,45,46 All the studies used the American College of Rheumatology criteria for RA diagnosis. For only the Pavkova et al. and the Lo et al. studies, the data was represented as allelic frequencies.32,34 Using the Newcastle-Ottawa scale, none of the studies were determined to contain significant study bias, but the Guseva et al. study9 and the Marinou et al. study9 could possible contain potential study bias. For the controls with respect to HWE, 5 studies for the −174 polymorphism,8,17,29,37,45 1 study for the −572 polymorphism,9 and 4 studies for the −597 polymorphism8,17,37,38 were determined to be not in agreement.
Characteristics of included studies.
| Study | Country (region) | RA criteria | Groupa | −174 G>Cb | −572 G>Cb | −597 G>Ab | NOSd |
|---|---|---|---|---|---|---|---|
| Ad’hiah 2018 | Iraq(Middle East) | ACR | Cases | –/–/– | 1/3/47 | –/–/– | 8 |
| Control | –/–/– | 0/0/45 | –/–/– | ||||
| Amr 2016 | Egypt(Middle East) | ACR | Cases | 44/46/9 | 26/64/9 | –/–/– | 7 |
| Control | 75/23/1 | 36/58/5 * | –/–/– | ||||
| Arman 2012 | Turkey(Middle East) | ACR | Cases | 95/62/21 | 143/31/4 | 97/59/22 | 8 |
| Control | 144/80/23 * | 194/52/1 | 133/86/28 * | ||||
| Dahlqvist 2002 | Sweden(Western Europe) | N/S | Cases | 83/121/51 | –/–/– | –/–/– | 7 |
| Control | 57/103/38 | –/–/– | –/–/– | ||||
| Dar 2017 | India(Middle East) | N/S | Cases | 20/8/6 | –/–/– | –/–/– | 7 |
| Control | 64/16/0 | –/–/– | –/–/– | ||||
| de Souza 2014 | Brazil(Latin America) | ACR | Cases | 49/14/1 | –/–/– | –/–/– | 8 |
| Control | 29/12/7 * | –/–/– | –/–/– | ||||
| Emonts 2011 | Netherlands(Western Europe) | ACR | Cases | 146/133/56 | –/–/– | –/–/– | 7 |
| Control | 146/231/82 | –/–/– | –/–/– | ||||
| Gaber 2103 | Egypt(Middle East) | ACR | Cases | 24/11/2 | –/–/– | –/–/– | 7 |
| Control | 9/1/0 | –/–/– | –/–/– | ||||
| Gomes-Silva 2018 | Brazil(Latin America) | ACR | Cases | 68/50/2 | –/–/– | –/–/– | 8 |
| Control | 72/43/5 | –/–/– | –/–/– | ||||
| Guseva 2016 | Russia(Eastern Europe) | ACR | Cases | 40/62/20 | –/–/– | –/–/– | 8 |
| Control | 85/162/54 | –/–/– | –/–/– | ||||
| Guseva 2018 | Russia(Eastern Europe) | N/S | Cases | 97/22/1 | –/–/– | –/–/– | 6 |
| Control | 165/3/0 | –/–/– | –/–/– | ||||
| Huang 2007 | China(Asia) | ACR | Cases | 97/22/1 | 4/15/101 | –/–/– | 8 |
| Control | 165/3/0 | 13/50/105 | –/–/– | ||||
| Julia 2007 | Spain(Western Europe) | ACR | Cases | –/–/– | –/–/– | 115/112/31 | 8 |
| Control | –/–/– | –/–/– | 87/66/28 * | ||||
| Kobayashi 2009 | Japan(Asia) | ACR | Cases | 137/0/0 | 10/45/82 | –/–/– | 8 |
| Control | 108/0/0 | 8/40/60 | –/–/– | ||||
| Li 2009 | China(Asia) | ACR | Cases | 48/11/1 | 52/6/2 | –/–/– | 8 |
| Control | 82/2/0 | 52/25/7 | –/–/– | ||||
| Li 2014a | China(Asia) | ACR | Cases | 247/7/2 | 23/125/108 | 249/5/2 | 8 |
| Control | 329/1/1 * | 41/152/138 | 330/0/1 * | ||||
| Li 2014b | China(Asia) | ACR | Cases | 613/124/15 | –/–/– | –/–/– | 8 |
| Control | 786/10/2 * | –/–/– | –/–/– | ||||
| Liu 2013 | China(Asia) | N/S | Cases | –/–/– | 12/36/39 | –/–/– | 8 |
| Control | –/–/– | 16/34/15 | –/–/– | ||||
| Lo 2008c | Taiwan(Asia) | ACR | Cases | –/–/– | 199: 87/311 | –/–/– | 8 |
| Control | –/–/– | 130: 33/227 | –/–/– | ||||
| Lu 2009 | China(Asia) | ACR | Cases | 120/27/1 | 5/18/125 | –/–/– | 8 |
| Control | 118/2/0 | 9/36/75 | –/–/– | ||||
| Marinou 2007 | UK(Western Europe) | ACR | Cases | 282/482/166 | –/–/– | –/–/– | 6 |
| Control | 154/226/80 | –/–/– | –/–/– | ||||
| Palomino-Morales 2009 | Spain(Western Europe) | ACR | Cases | 127/144/40 | –/–/– | –/–/– | 8 |
| Control | 103/94/29 | –/–/– | –/–/– | ||||
| Panoulas 2009 | UK(Western Europe) | ACR | Cases | 135/176/72 | –/–/– | –/–/– | 7 |
| Control | 148/211/63 | –/–/– | –/–/– | ||||
| Pascual 2000 | Spain(Western Europe) | ACR | Cases | 75/72/16 | –/–/– | –/–/– | 7 |
| Control | 73/66/18 | –/–/– | –/–/– | ||||
| Pavkova 2014c | Czech Republic(Eastern Europe) | ACR | Cases | 144: 159/129 | –/–/– | –/–/– | 8 |
| Control | 200: 232/168 | –/–/– | –/–/– | ||||
| Pawlik 2005 | Poland(Eastern Europe) | ACR | Cases | 26/53/19 | –/–/– | –/–/– | 8 |
| Control | 25/58/22 | –/–/– | –/–/– | ||||
| Raafat Hamed 2018 | Egypt(Middle East) | ACR | Cases | 15/8/2 | –/–/– | –/–/– | 7 |
| Control | 25/0/0 | –/–/– | –/–/– | ||||
| Schotte 2015 | Germany(Western Europe) | ACR | Cases | 17/24/9 | 46/4/0 | 17/24/9 | 7 |
| Control | 30/36/25 * | 85/6/0 | 30/36/25 * | ||||
| Shafia 2014 | India(Middle East) | ACR | Cases | 122/27/1 | –/–/– | –/–/– | 8 |
| Control | 167/30/3 | –/–/– | –/–/– | ||||
| Trajkov 2009 | Macedonia(Western Europe) | ACR | Cases | 38/30/16 | –/–/– | 38/33/13 | 7 |
| Control | 144/132/25 | –/–/– | 153/123/25 | ||||
| Wielinska 2018 | Poland(Eastern Europe) | ACR | Cases | 40/65/25 | –/–/– | –/–/– | 7 |
| Control | 37/53/22 | –/–/– | –/–/– | ||||
| You 2013 | China(Asia) | ACR | Cases | 431/21/0 | 38/191/222 | 418/31/3 | 8 |
| Control | 357/16/0 | 39/166/168 | 359/14/0 | ||||
| Zavaleta-Muñiz 2013 | Mexico(Latin America) | ACR | Cases | 106/30/1 | 74/58/5 | –/–/– | 8 |
| Control | 80/20/2 | 62/37/3 | –/–/– |
Abbreviations: ACR: American College of Rheumatology; N/S: not specified; NOS: New Castle-Ottawa Scale.
For the controls. HWE was calculated using ψ2-test. p<0.05 is not in agreement with HWE and indicated with an *.
The ratios are given as Wild type, heterozgotes, and homozygote mutant. For the −174 polymorphism, the G-allele is considered the Wild-type and C-allele as the mutant. For the −572 polymorphism, the G-allele is considered the Wild-type and C-allele as the mutant. For the −597 polymorphism, the G-allele is considered the Wild-type and A-allele as the mutant.
The ORs and 95%CIs were calculated for each study for all 5 genetic models (all Forest plots are available as supplement information). Using the selected studies, for the −174 polymorphism, there was significant heterogeneity for all the genetic models; therefore, the random-effects model was utilized (Table 2). When the studies were pooled together, the heterozygous (OR: 1.47, 95%CI: 1.13–1.91, p=0.004), dominant (OR: 1.50, 95%CI: 1.15–1.96, p=0.003), and allelic (OR: 1.38, 95%CI: 1.13–1.68, p=0.001) genetic models demonstrated a significant association. Removing one study did not change the association for any of the genetic models.
The association between IL-6 promoter polymorphism and the risk of developing Rheumatoid Arthritis.
| Heterogeneityb | Associationc | Publication Biasd | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Genetic model | Analysis | Na | Qp-value | I2 (%) | Model | OR | 95% CI | p-Value | Begg | Egger |
| -174 G>C | ||||||||||
| Heterozygous | Overall | 27 | <0.01 | 82 | Random | 1.47 | 1.13–1.91 | 0.004* | Tau=0.39, p<0.01* | Bias=2.78, p<0.01* |
| Homozygous | Overall | 26 | <0.01 | 47 | Random | 1.13 | 0.88–1.44 | 0.337 | Tau=0.12, p=0.39 | Bias=0.67, p=0.14 |
| Dominant | Overall | 27 | <0.01 | 84 | Random | 1.50 | 1.15–1.96 | 0.003* | Tau=0.31, p=0.02* | Bias=2.87, p<0.01* |
| Recessive | Overall | 26 | 0.03 | 38 | Random | 1.12 | 0.92–1.37 | 0.270 | Tau=0.09, p=0.49 | Bias=0.49, p=0.23 |
| Allelic | Overall | 28 | <0.01 | 85 | Random | 1.38 | 1.13–1.68 | 0.001* | Tau=0.35, p<0.01* | Bias=2.73, p<0.01* |
| -572 G>C | ||||||||||
| Heterozygous | Overall | 11 | 0.17 | 29 | Fixed | 1.09 | 0.88–1.35 | 0.420 | Tau=-0.09, p=0.70 | Bias=-0.94, p=0.39 |
| Homozygous | Overall | 11 | 0.21 | 24 | Fixed | 1.60 | 1.21–2.12 | 0.001* | Tau=-0.13, p=0.59 | Bias=0.21, p=0.80 |
| Dominant | Overall | 12 | 0.04 | 46 | Random | 1.21 | 0.90–1.63 | 0.208 | Tau=0.09, p=0.68 | Bias=-0.11, p=0.92 |
| Recessive | Overall | 11 | <0.01 | 68 | Random | 1.61 | 1.11–2.33 | 0.012* | Tau=-0.09, p=0.70 | Bias=0.56, p=0.59 |
| Allelic | Overall | 13 | <0.01 | 79 | Random | 1.18 | 0.90–1.54 | 0.246 | Tau=0.03, p=0.90 | Bias=-0.46, p=0.75 |
| -597 G>A | ||||||||||
| Heterozygous | Overall | 6 | 0.28 | 21 | Fixed | 1.20 | 0.96–1.51 | 0.115 | Tau=0.47, p=0.19 | Bias=1.91, p=0.09 |
| Homozygous | Overall | 6 | 0.25 | 24 | Fixed | 1.10 | 0.79–1.55 | 0.575 | Tau=0.33, p=0.34 | Bias=1.13, p=0.32 |
| Dominant | Overall | 6 | 0.15 | 38 | Fixed | 1.19 | 0.97–1.48 | 0.100 | Tau=0.47, p=0.19 | Bias=2.21, p=0.09 |
| Recessive | Overall | 6 | 0.14 | 40 | Fixed | 1.03 | 0.75–1.42 | 0.854 | Tau=0.33, p=0.34 | Bias=1.21, p=0.35 |
| Allelic | Overall | 6 | 0.02 | 61 | Random | 1.21 | 0.90–1.61 | 0.205 | Tau=0.47, p=0.19 | Bias=2.68, p=0.10 |
Abbreviations: OR: Odds ratio; 95%CI: 95% confidence interval; and I2: Inconsistency Index.
Publication bias was assessed by examining the funnel plot. The funnel plots demonstrated no significant asymmetry and the shape of the funnel plots suggested no evidence of publication bias (see Supplement information), even though some over-dispersion can be seen. No correlation was determined by the Begg-Mazumdar's test or bias by Egger's test for the homozygous and recessive genetic models (Table 2); however, there was the presence of publication bias for the heterozygous, dominant, and allelic genetic models (Begg–Mazumdar's test: p<0.01, Egger's test: p<0.01).
When the cohort was stratified by geographic region, for the Asian population, the −174 polymorphism did demonstrate a strong association for all the genetic models (Table 3). For Middle East countries, the −174 polymorphism showed an association for the heterozygous (OR: 2.30, 95%CI: 1.10–4.81, p=0.028), dominant (OR: 2.27, 95%CI: 1.22–4.22, p=0.010), and allelic (OR: 2.29, 95%CI: 1.24–4.23, p=0.008) genetic models. Interestingly, for Latin America, the −174 polymorphism demonstrated a protective benefit for the homozygous (OR: 0.26, 95%CI: 0.08–0.82, p=0.022) and recessive (OR: 0.25, 95%CI: 0.08–0.80, p=0.019) genetic models.
The association of the −174 G>C polymorphism on the development of Rheumatoid Arthritis, stratified by ethnicity.
| Heterogeneityb | Associationc | |||||||
|---|---|---|---|---|---|---|---|---|
| Genetic model | Analysis | Na | Qp-value | I2 (%) | Model | OR | 95% CI | p-Value |
| Asian | ||||||||
| Heterozygous | 6 | <0.01 | 86 | Random | 7.57 | 2.28–25.14 | 0.001* | |
| Homozygous | 5 | 0.91 | 0 | Fixed | 5.84 | 2.06–16.56 | 0.001* | |
| Dominant | 6 | <0.01 | 87 | Random | 7.21 | 2.30–22.63 | 0.001* | |
| Recessive | 5 | 0.92 | 0 | Fixed | 5.04 | 1.78–14.28 | 0.002* | |
| Allelic | 6 | <0.01 | 86 | Random | 6.60 | 2.26–19.28 | 0.001* | |
| Middle East | ||||||||
| Heterozygous | 5 | 0.02 | 66 | Random | 2.30 | 1.10–4.81 | 0.028* | |
| Homozygous | 6 | 0.05 | 56 | Random | 3.42 | 0.94–12.42 | 0.062 | |
| Dominant | 6 | <0.01 | 72 | Random | 2.27 | 1.22–4.22 | 0.010* | |
| Recessive | 6 | 0.09 | 48 | Random | 2.69 | 0.83–8.66 | 0.098 | |
| Allelic | 6 | <0.01 | 79 | Random | 2.29 | 1.24–4.23 | 0.008* | |
| East Europe | ||||||||
| Heterozygous | 6 | 0.93 | 0 | Fixed | 0.91 | 0.72–1.14 | 0.393 | |
| Homozygous | 5 | 0.18 | 36 | Fixed | 1.09 | 0.81–1.49 | 0.562 | |
| Dominant | 5 | 0.85 | 0 | Fixed | 0.94 | 0.75–1.17 | 0.581 | |
| Recessive | 5 | 0.14 | 43 | Fixed | 1.15 | 0.88–1.51 | 0.307 | |
| Allelic | 6 | 0.56 | 0 | Fixed | 1.03 | 0.90–1.18 | 0.650 | |
| West Europe | ||||||||
| Heterozygous | 7 | 0.01 | 63 | Random | 0.96 | 0.76–1.20 | 0.689 | |
| Homozygous | 7 | 0.40 | 4 | Fixed | 0.99 | 0.82–1.18 | 0.876 | |
| Dominant | 7 | 0.03 | 58 | Random | 0.95 | 0.77–1.16 | 0.606 | |
| Recessive | 7 | 0.66 | 0 | Fixed | 1.03 | 0.87–1.21 | 0.759 | |
| Allelic | 7 | 0.14 | 37 | Fixed | 0.99 | 0.90–1.08 | 0.732 | |
| Latin | ||||||||
| Heterozygous | 3 | 0.55 | 0 | Fixed | 1.09 | 0.75–1.57 | 0.664 | |
| Homozygous | 3 | 0.48 | 0 | Fixed | 0.26 | 0.08–0.82 | 0.022* | |
| Dominant | 3 | 0.17 | 43 | Fixed | 0.94 | 0.66–0.35 | 0.750 | |
| Recessive | 3 | 0.55 | 0 | Fixed | 0.25 | 0.08–0.80 | 0.019* | |
| Allelic | 3 | <0.05 | 67 | Random | 0.77 | 0.44–1.34 | 0.353 | |
Abbreviations: OR: Odds ratio; 95%CI: 95% confidence interval; and I2: Inconsistency Index.
For the −572 polymorphism, the homozygous and heterozygous genetic models did not present with a significant level of heterogeneity and were analyzed using fixed-effects, whereas the rest of the genetic models were analyzed with random-effects. Only the homozygous (OR: 1.60, 95%CI: 1.21–2.12, p=0.001) and recessive (OR: 1.61, 95%CI: 1.11–2.33, p=0.012) genetic models demonstrated a positive association between the polymorphism and the risk of developing RA (Table 2). Removing the Li et al. study9 resulted in a significant positive association for the dominant genetic model (OR: 1.32, 95%CI: 1.08–1.63, p=0.008), none of the other studies had any effect. For the remaining genetic model, the results were resilient to any change from removing one study. No publication bias was determined by examining the Funnel plot, Egger's bias test, or by Begg–Mazumdar's correlation test.
When stratified by geographic region, it appears that the association was due to the inclusion of studies from the Asian population (Table 4). For the Asian population alone, there was an association between the −572 polymorphism and the development of RA for the homozygous (OR: 1.56, 95%CI: 1.16–2.09, p=0.004) and recessive genetic models (OR: 1.63, 95%CI: 1.08–2.45, p=0.020). No association was observed for the Middle East region.
The association of the −572 G>C polymorphism on the development of Rheumatoid Arthritis, stratified by ethnicity.
| Heterogeneityb | Associationc | |||||||
|---|---|---|---|---|---|---|---|---|
| Genetic model | Analysis | Na | Q p-value | I2 (%) | Model | OR | 95% CI | p-Value |
| Asian | ||||||||
| Heterozygous | 7 | 0.09 | 46 | Random | 0.97 | 0.63–1.49 | 0.883 | |
| Homozygous | 7 | 0.10 | 43 | Fixed | 1.56 | 1.16–2.09 | 0.004* | |
| Dominant | 7 | <0.01 | 66 | Random | 1.23 | 0.74–2.05 | 0.416 | |
| Recessive | 7 | <0.01 | 78 | Random | 1.63 | 1.08–2.45 | 0.020* | |
| Allelic | 8 | <0.01 | 86 | Random | 1.20 | 0.82–1.74 | 0.351 | |
| Middle East | ||||||||
| Heterozygous | 2 | 0.12 | 60 | Random | 1.08 | 0.58–2.01 | 0.806 | |
| Homozygous | 3 | 0.39 | 0 | Fixed | 2.42 | 0.89–6.61 | 0.084 | |
| Dominant | 3 | 0.27 | 24 | Fixed | 1.10 | 0.76–1.60 | 0.604 | |
| Recessive | 3 | 0.11 | 54 | Fixed | 1.73 | 0.67–4.48 | 0.260 | |
| Allelic | 3 | 0.15 | 46 | Fixed | 1.15 | 0.85–1.54 | 0.367 | |
Abbreviations: OR: Odds ratio; 95%CI: 95% confidence interval; and I2: Inconsistency Index.
For the −597 polymorphism, all the genetic models were analyzed with the fixed-effects model, except the allelic genetic model. When the studies were pooled together, there was no effect for any of the genetic models (Table 2). However, removing the Arman et al. study,8 a positive association for the heterozygous (OR: 1.33, 95%CI: 1.01–1.74, p=0.040) and dominant (OR: 1.30, 95%CI: 1.01–1.68, p=0.040) genetic models was observed. No publication bias was determined by examining the Funnel plot, Egger's bias test, or by Begg–Mazumdar's correlation test. No sub-analysis could be performed for the −597 polymorphism, due to the few studies.
DiscussionSome studies have shown that IL-6 serum levels are associated with the development of RA14; moreover, it has been postulated that the IL6 −174, −572, and −597 polymorphisms are associated with RA development. Here, we show that the −597 polymorphism does not promote RA development; however, the −174 and the −572 polymorphisms do indeed increase the risk of developing RA, especially in Asian and Middle East countries. Unexpectedly, the −174 polymorphism showed a protective effect for Latin American countries.
Previous meta-analyses have shown that the −174 polymorphism does augment the risk of developing RA, especially for Asian populations and not others.15–17,47 Our results also confirm this; however, we also found an effect for the −174 polymorphism with countries that are from the Middle East. When the genotype distributions were examined, it appeared that these Middle East and Asian populations had the lowest minor allele frequencies (10% and 1%, respectively), which were different from European populations (∼40%). Since the GG genotype is associated with higher serum IL-6 and incremental decreases in serum IL-6 were associated with each additional C-allele,48 this would suggest that a large portion of the Europe population would have lower IL-6 serum levels. However, serum IL-6 are also affected by many confounding factors. Serum IL-6 was shown to be affected by age, circadian rhythm, and stress as well as overweight or obesity patients present with elevated serum IL-6 when compared to normal weight subjects.49–52 Lastly, it is demonstrated that diets with phytoestrogens decrease serum IL-6,53 whereas, diet with increase carbohydrates or monounsaturated fat augment serum IL-6.54,55 Therefore lifestyle, diet, or other factors could significantly affect serum IL-6 level and mitigate the −174 polymorphism's effect associated with RA development.
There are many studies that show differences in the IL-6 serum levels is ethnic dependent. In African Americans, the production of IL-6 was higher compared to Cuban Americans.47 Whereas, in Hispanics, especially Mexicans, IL-6 production was demonstrated to be lower than other ethnicities.51 However, promoter polymorphisms have shown to increase the production of IL-6,14 but the prevalence of these polymorphisms varying significantly from region to region. For example, Gao et al. reported the prevalence of −174 polymorphism was less frequent in Asian Indian, Afro-Caribbean, Afro-American, and Asians.13
Here, we show that, for Latinos, the homozygous mutant of the −174 polymorphism was associated with a protective effect. The mechanism for this result remains elusive. Even though the −174 polymorphism was shown to increase IL-6 production, with a lower basal level, it is possible that the increases are insufficient to augment the risk of developing RA. Moreover, the quality of life and lifestyle in Latin American, which has not fully adopted the Western lifestyle, these confounding factors can mitigate the effect elevated IL-6 causes in developing RA. In support of this, it has been shown that the type of diet affects the production of IL-6 from muscle cells, which indirectly affects immune cell penetration of muscle tissue, promoting prolong IL-6 release.56 Diets, in which more carbohydrates are consumed, does mitigate the secondary IL-6 peak, caused by immune cells, after moderate exercise.57 Moreover, the more active the subject's lifestyle, the lower IL-6 production.58,59 In Latin American countries, it is possible that the lifestyle and quality of life does mitigate the effect the −174 polymorphism has on the development of RA in Latin American. This does posit that some factors such as lifestyle or diet in the presence of elevated IL-6 levels does sensitize a subject to an anti-inflammatory state, leading to the increased release of IL-6 and the development of RA; however, more studies are required to elucidated if the protective effect against RA development is connected to these confounding factors. In a review by Saavedra Ramirez et al., they eloquently explain the enigma of IL-6 in disease pathogenesis and present the possible switch between pro- and anti-inflammatory functions of IL-6.60
Three previous meta-analyses did examine for any associations between the −572 and −597 polymorphisms with RA development-one conducted in 2012, which used 2 and 1 studies, respectively, one in 2014, which used 6 and 2 studies, respectively, and one in 2019 that used 6 studies for the −572 polymorphism.17,47,61 Since then, numerous studies have examined the association and, in our study using 6 studies, we found no association between the −597 polymorphism and the development of RA. However, removing the Arman et al. study did indicate that an association could exist. Therefore, more research is required to deduce the association between RA and the −597 polymorphism.
For the −572 polymorphism, using 13 studies, we found that the homozygous mutant does increase the risk of developing RA; however, this result appears to be due the Asian population. This is in disagreement with the Li et al. meta-analysis17; however, we would say our study is in agreement with the Dar et al. meta-analysis,15 but their result was based-off of the Huang et al. study, which is included here. Recently, in 2019, a systematic review by Zhang et al., which solely focused on the −572 polymorphism, did determined that the polymorphism is associated with the risk of RA, specially the GG genotype.62 We believe that their study presents with similar results, but our study appears to be more inclusive. Here, we have 13 studies, whereas Zhang et al. only used 6 studies, of which all were included in our study, suggesting a possibility of publication/selection bias. Indeed, when we compared the results, we found that for their dominant model (our study's recessive model), the result did not concur, which highly suggests the presence of selection bias. Nevertheless, as we postulated, any effect observed was due to the Asian population and Zhang et al. confirmed our observation.62 However, it must be noted that Zhang et al.’s Asian population consisted of 3 studies, whereas our Asian population consisted of 8 studies. This would suggest that our results are more stable than Zhang et al. It could be expected that the Middle East region should have presented with an association; however, with the few studies included here, we could not determine this result.62
With our meta-analysis, some models presented with significant heterogeneity. We believe the cause could be due to the cases and controls, which were not matched by sex (majority of cases were women), age, or type of RA. In a previous study by Donn et al., in juvenile idiopathic arthritis, there was a significant difference in the ratio between systemic onset and enthesitis-related,63 thus indicating that the type of RA could be affected differently. Another cause could be the subject's ethnicity. Even though we performed an ethnic sub-analysis, genetic differences within a country can vary significantly, as indicated in China,64,65 Mexico,66 Brazil,67,68 and India.69 The diagnostic criteria used to identify or categorize RA could also act as a source of heterogeneity.
This study has a few limitations. First, as mentioned above, due to the large genetic variation within a country, more studies are required to determine specific effects, such as the protective benefit our results indicated for Latin Americans. Second, our results can only focus on overall risk of developing RA, and stage-specific effects (low, moderate, high or remission) cannot be determined. Third, we calculated crude ORs that were not adjusted.
Here, we show that the −174 polymorphism increased the risk of developing RA for Asians and Middle East populations; interestingly, there was a protective effect for Latinos. As for the −572 polymorphism, only the Asian population was associated with an increased risk of developing RA. No affect was observed for the −597 polymorphism.
Conflicts of interestsThe authors declare that they have no conflicts of interests to report.
We would like to express their gratitude to Mtro. Ricardo Villegas Tovar, Coordinator of Scientific Production and International Visibility, Benemérita Universidad Autónoma de Puebla. Moreover, we would like to thank Dr. Solbritt Rantapää Dahlqvist, Dr. Irina Anatolyevna Guseva, Dr. Anthony G Wilson, and Dr. Heiko Schotte for their prompt replies to our emails and the sharing of the data to allow a more conclusive analysis between IL6 promoter polymorphisms and the development of RA. This study was supported by grants from the Vicerrectoría de Investigación of Benemérita Universidad Autónoma de Puebla (10051909-VIEP2018 to MEGM and 100170644-VIEP2018 to RPF. 100493499-VIEP2018 to ETR).